# --- Find MLE by running GD to convergence ---
a_mle, b_mle = 0.5, 0.3
for _ in range(100000):
phi0 = np.exp(-0.5 * X2**2) / np.sqrt(2 * np.pi)
phi_b = np.exp(-0.5 * (X2 - b_mle)**2) / np.sqrt(2 * np.pi)
px = (1 - a_mle) * phi0 + a_mle * phi_b + 1e-15
grad_a = np.mean((phi0 - phi_b) / px)
grad_b = np.mean(-a_mle * (X2 - b_mle) * phi_b / px)
a_mle -= 0.001 * grad_a
b_mle -= 0.001 * grad_b
a_mle = np.clip(a_mle, 0.01, 0.99)
# --- Empirical SGD trajectory statistics ---
traj = cached_sgd(sgd_model2, X2, 2.0*batch_size2/n2, batch_size2,
n_steps=300000, burn_in=10000, seed=123)
sgd_mean = traj.mean(axis=0)
emp_cov = np.cov(traj.T)
evals_emp, evecs_emp = np.linalg.eigh(emp_cov)
print(f"MLE: a* = {a_mle:.4f}, b* = {b_mle:.4f}")
print(f"SGD traj. mean: a = {sgd_mean[0]:.4f}, b = {sgd_mean[1]:.4f}")
print(f"Shift from MLE: Δa = {sgd_mean[0]-a_mle:.4f}, Δb = {sgd_mean[1]-b_mle:.4f}\n")
# Helper: compute H and C at a given point
eps = 1e-5
def nll(a_val, b_val):
phi0_ = np.exp(-0.5 * X2**2) / np.sqrt(2 * np.pi)
phi_b_ = np.exp(-0.5 * (X2 - b_val)**2) / np.sqrt(2 * np.pi)
return -np.mean(np.log((1 - a_val) * phi0_ + a_val * phi_b_ + 1e-15))
def compute_H_C(a0, b0):
f0 = nll(a0, b0)
H_ = np.array([
[(nll(a0+eps,b0) - 2*f0 + nll(a0-eps,b0)) / eps**2,
(nll(a0+eps,b0+eps) - nll(a0+eps,b0-eps)
- nll(a0-eps,b0+eps) + nll(a0-eps,b0-eps)) / (4*eps**2)],
[0, (nll(a0,b0+eps) - 2*f0 + nll(a0,b0-eps)) / eps**2]])
H_[1,0] = H_[0,1]
phi0_ = np.exp(-0.5 * X2**2) / np.sqrt(2*np.pi)
phi_b_ = np.exp(-0.5 * (X2 - b0)**2) / np.sqrt(2*np.pi)
px_ = (1 - a0)*phi0_ + a0*phi_b_ + 1e-15
g = np.column_stack([(phi0_ - phi_b_)/px_, -a0*(X2 - b0)*phi_b_/px_])
C_ = (g.T @ g)/len(X2)
C_ -= g.mean(0,keepdims=True).T @ g.mean(0,keepdims=True)
return H_, C_
B_batch = 10
# OU prediction at MLE
H_mle, C_mle = compute_H_C(a_mle, b_mle)
Sigma_mle = np.linalg.inv(H_mle) @ C_mle / (2*B_batch)
evals_mle, evecs_mle = np.linalg.eigh(Sigma_mle)
# OU prediction at SGD mean
H_sgd, C_sgd = compute_H_C(sgd_mean[0], sgd_mean[1])
Sigma_sgd = np.linalg.inv(H_sgd) @ C_sgd / (2*B_batch)
evals_sgd, evecs_sgd = np.linalg.eigh(Sigma_sgd)
# Report
def report(label, evals, evecs):
angle = np.degrees(np.arctan2(evecs[1,1], evecs[0,1]))
print(f"{label}:")
print(f" eigenvalues = {evals[0]:.6f}, {evals[1]:.6f} (ratio {evals[1]/evals[0]:.1f}:1)")
print(f" major axis angle = {angle:.1f}°")
report("Empirical covariance", evals_emp, evecs_emp)
report("OU prediction @ MLE", evals_mle, evecs_mle)
report("OU prediction @ SGD mean", evals_sgd, evecs_sgd)
# --- Visualise ---
from matplotlib.patches import Ellipse
fig, ax = plt.subplots(figsize=(7, 5))
# Posterior contours
ax.contour(A_grid2, B_grid2, posterior2 / np.max(posterior2),
levels=[0.1, 0.3, 0.5, 0.7, 0.9],
colors='royalblue', linewidths=1.5, alpha=0.6)
# SGD density
hist, _, _ = np.histogram2d(traj[:, 0], traj[:, 1], bins=80,
range=[[0.01, 0.99], [-0.5, 1.0]])
hist = gaussian_filter(hist.T, sigma=1.5)
hist /= np.max(hist)
ax.imshow(hist, origin='lower', extent=[0.01, 0.99, -0.5, 1.0],
aspect='auto', cmap='Oranges', alpha=0.7)
# Posterior mean from grid
da2 = a_grid2[1] - a_grid2[0]
db2 = b_grid2[1] - b_grid2[0]
norm = np.sum(posterior2) * da2 * db2
post_mean_a = np.sum(A_grid2 * posterior2) * da2 * db2 / norm
post_mean_b = np.sum(B_grid2 * posterior2) * da2 * db2 / norm
print(f"\nPosterior mean: a = {post_mean_a:.4f}, b = {post_mean_b:.4f}")
# Mark MLE, posterior mean, and SGD mean
ax.plot(a_mle, b_mle, 'k+', markersize=12, markeredgewidth=2, label='MLE')
ax.plot(post_mean_a, post_mean_b, 'D', color='royalblue', markersize=8,
markeredgecolor='white', markeredgewidth=1, label='Posterior mean')
ax.plot(sgd_mean[0], sgd_mean[1], 'rx', markersize=10, markeredgewidth=2,
label='SGD mean')
# Draw eigenvector arrows
scale = 0.15
# At MLE: H small-eigenvalue direction (posterior ridge) in blue
evals_Hm, evecs_Hm = np.linalg.eigh(H_mle)
v_post = evecs_Hm[:, 0] # small eigenvalue = flat/ridge direction
for sign in [1, -1]:
ax.annotate('', xy=(a_mle + sign*scale*v_post[0], b_mle + sign*scale*v_post[1]),
xytext=(a_mle, b_mle),
arrowprops=dict(arrowstyle='->', color='blue', lw=2.5))
# At MLE: OU-predicted Σ large-eigenvalue direction in orange
v_ou = evecs_mle[:, 1] # large eigenvalue = predicted SGD stretch
for sign in [1, -1]:
ax.annotate('', xy=(a_mle + sign*scale*v_ou[0], b_mle + sign*scale*v_ou[1]),
xytext=(a_mle, b_mle),
arrowprops=dict(arrowstyle='->', color='darkorange', lw=2.5))
# At SGD mean: empirical major axis in green
v_emp = evecs_emp[:, 1]
for sign in [1, -1]:
ax.annotate('', xy=(sgd_mean[0] + sign*scale*v_emp[0], sgd_mean[1] + sign*scale*v_emp[1]),
xytext=(sgd_mean[0], sgd_mean[1]),
arrowprops=dict(arrowstyle='->', color='green', lw=2.5))
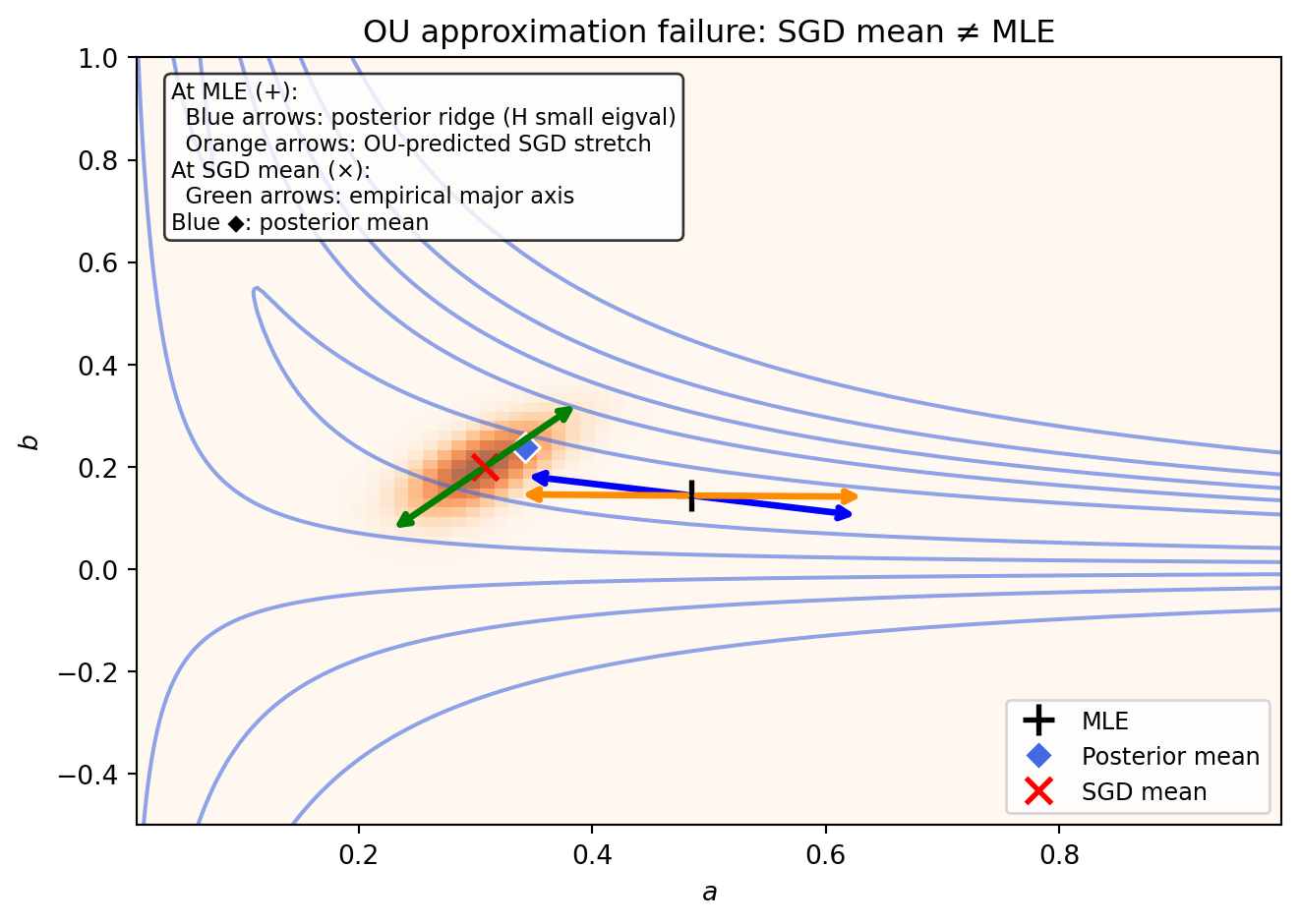
ax.text(0.03, 0.97, 'At MLE (+):\n'
' Blue arrows: posterior ridge (H small eigval)\n'
' Orange arrows: OU-predicted SGD stretch\n'
'At SGD mean (×):\n'
' Green arrows: empirical major axis\n'
'Blue ◆: posterior mean',
transform=ax.transAxes, fontsize=8.5, va='top',
bbox=dict(boxstyle='round', facecolor='white', alpha=0.8))
ax.legend(loc='lower right', fontsize=9)
ax.set_xlabel('$a$')
ax.set_ylabel('$b$')
ax.set_title('OU approximation failure: SGD mean ≠ MLE')
plt.tight_layout()
plt.show()